3D-Fast optical interface for rapid volumetric neural sensing and modulation
(collaboration with Drs. Cristin Welle, CU Anschutz Neurosurgery, Emily Gibson, CU Anschutz Bioengineering, Doug Shepherd, CU Anschutz Pharmacology and Ioannis Kymissis, Columbia University).
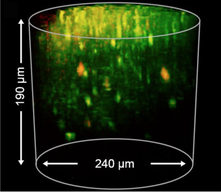
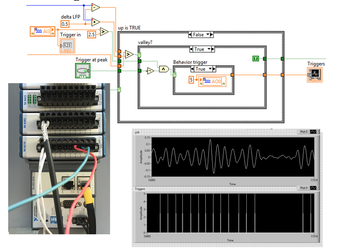
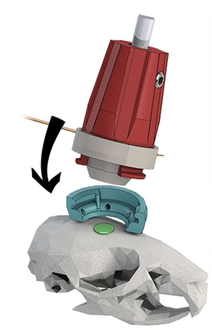
We propose to develop an optical device (‘3D-FAST’) that allows for rapid, realtime volumetric neuronal recording of calcium and voltage transients and precise optical stimulation. By pairing miniature arrays of micropatterned LED emitters with the axial focusing capabilities of electrowetting lens technologies, we will achieve duplex recording and stimulation of many thousands of neurons.
Welle (PI) and Gibson (PI). Funded by NIH BRAIN Initiative R01 NS123665 Welle (PI) and Gibson (PI).
(collaboration with Drs. Cristin Welle, CU Anschutz Neurosurgery, Emily Gibson, CU Anschutz Bioengineering, Doug Shepherd, CU Anschutz Pharmacology and Ioannis Kymissis, Columbia University).
We propose to develop an optical device (‘3D-FAST’) that allows for rapid, realtime volumetric neuronal recording of calcium and voltage transients and precise optical stimulation. By pairing miniature arrays of micropatterned LED emitters with the axial focusing capabilities of electrowetting lens technologies, we will achieve duplex recording and stimulation of many thousands of neurons.
Welle (PI) and Gibson (PI). Funded by NIH BRAIN Initiative R01 NS123665 Welle (PI) and Gibson (PI).
|
Cortical circuit dysfunction in fragile x syndrome
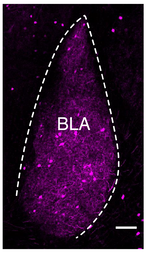
(collaboration with Dr. Molly Huntsman). The experiments define in an animal model system for Fragile X Syndrome mechanisms resulting in faulty activation of a small yet powerful population of inhibitory interneurons, and identify modes of corrective restoration through pharmacologic and genetic approaches. Funded by NINDS R01NS095311, Huntsman (PI), Restrepo (Co-PI) |
|
Development of a Bidirectional Optogenetic Minimally Invasive Peripheral Nerve Interface with single axon read-in & read-out specificity(Collaboration with Drs. Richard Weir, John Caldwell and Emily Gibson)
Our goal for this project is to develop a novel compact Optogenetic based Optical Probe capable of optically neuromodulating individual afferent and/or efferent axons within nerves of the parasympathetic, or peripheral, nervous system. We seek to read-in or read-out from these nerves with the goal of modulating the organs or brain circuits innervated by them. Funded by NIH SPARC, Caldwell (PI), Gibson(PI), Weir (PI). |
Scalable 3D molecular imaging and data analysis for cell census generation
(collaboration with Drs. Douglas Shepherd and Steve Presse, Arizona State University).
This project is a collaboration across two universities and multiple scientific disciplines to develop new scalable 3D molecular imaging and analysis approaches for cell type identification within human brain tissue. We will focus our efforts on the olfactory system, comprising the olfactory epithelium (OE) and the olfactory bulb (OB). Our long term goals are to provide the community with new “physics-first” methods that improve scalability, rigor, and 3D measurements for cell census creation and create the first spatial map of connections between the OE and OB.
Shepherd (PI) and Presse (PI). Funded by NIH BRAIN Initiative R01 MH128867 Shepherd (PI) and Presse (PI).
(collaboration with Drs. Douglas Shepherd and Steve Presse, Arizona State University).
This project is a collaboration across two universities and multiple scientific disciplines to develop new scalable 3D molecular imaging and analysis approaches for cell type identification within human brain tissue. We will focus our efforts on the olfactory system, comprising the olfactory epithelium (OE) and the olfactory bulb (OB). Our long term goals are to provide the community with new “physics-first” methods that improve scalability, rigor, and 3D measurements for cell census creation and create the first spatial map of connections between the OE and OB.
Shepherd (PI) and Presse (PI). Funded by NIH BRAIN Initiative R01 MH128867 Shepherd (PI) and Presse (PI).
|
Collaborative Research: NCS-FR: Shedding light on brain circuits mediating navigation of the odor plume in a natural environment
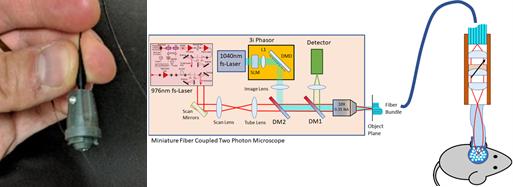
Members of our OPeN interdisciplinary team developed a novel two photon fiber-coupled microscope for 3D imaging of brain activity in the freely moving mouse under partial funding from an NSF NCS:FOUNDATIONS grant and generated and quantified realistic odor environments in the laboratory to explore algorithms used for odor-guided navigation under funding by an NSF Ideas Lab “Cracking the Olfactory Code”. In this NCS:FRONTIERS project we leverage the extensive expertise and achievements of the team to crack the circuit basis for odor plume navigation. We will develop a low-weight, miniature 3-photon fiber coupled microscope (3P-FCM) to record neuronal activity simultaneously in one brain area in two planes of view. In addition, OPeN will develop a portable photoionization (PID) sensor to detect the odorant concentration at the nostril as the animal navigates the odor plume. Members of the OPeN team will record neural activity in the hippocampus and cerebellum of animals navigating the odor plume and will develop a Bayesian analysis method to decode odor plume navigation from neural activity. This multidisciplinary approach will result in understanding of the brain mechanisms of odor plume navigation. |
Having fun following up on Eichenbaum’s episodic memory for odorants in CA1 |
1. Gire DH, Whitesell JD, Doucette W, Restrepo D. Information for decision-making and stimulus identification is multiplexed in sensory cortex. Nat Neurosci. 2013;16(8):991-3. Epub 2013/06/25. doi: 10.1038/nn.3432. PubMed PMID: 23792942; PMCID: PMC3725200.
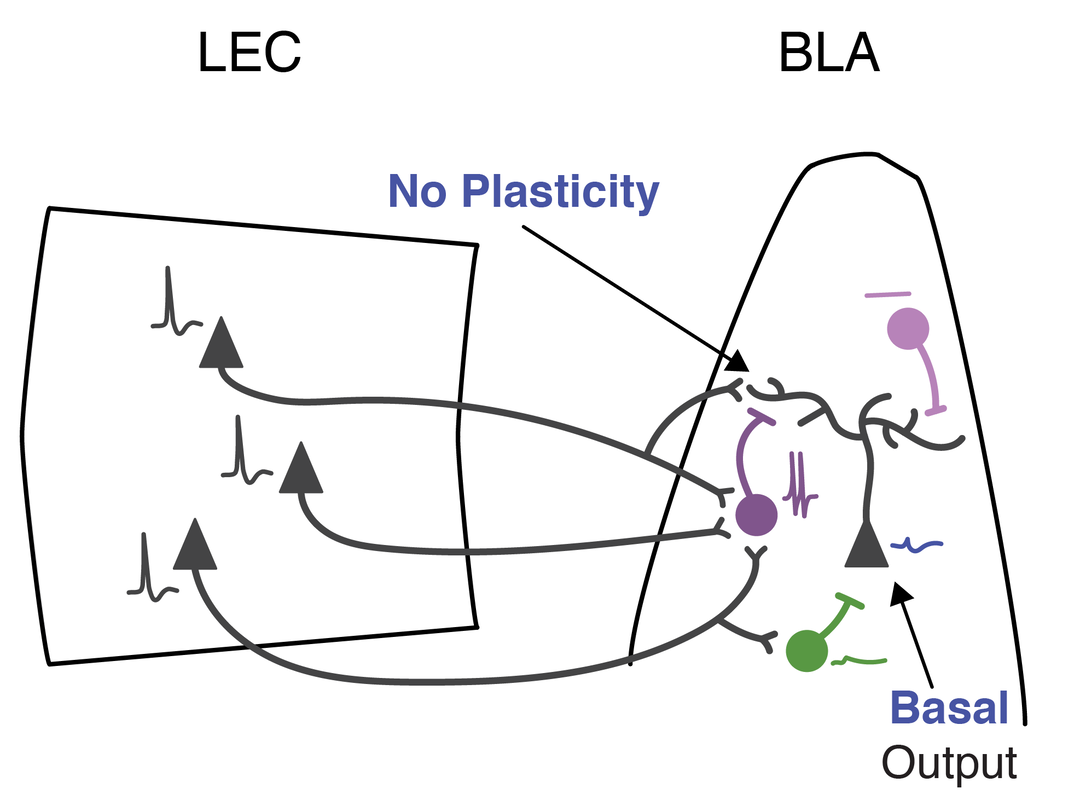
2. Guthman EM, Ma M, Chu P, Baca SM, Restrepo D, Huntsman MM. Cell type specific control of basolateral amygdala plasticity via feedforward inhibition. bioRxiv. 2018. doi: https://doi.org/10.1101/348524.
3. Gould EA, Busquet N, Shepherd D, Dietz RM, Herson PS, Simoes de Souza FM, Li A, George NM, Restrepo D, Macklin WB. Mild myelin disruption elicits early alteration in behavior and proliferation in the subventricular zone. Elife. 2018;7. Epub 2018/02/14. doi: 10.7554/eLife.34783. PubMed PMID: 29436368; PMCID: PMC5828668.
4. Ozbay BN, Futia GL, Ma M, Bright VM, Gopinath JT, Hughes EG, Restrepo D, Gibson EA. Three dimensional two-photon brain imaging in freely moving mice using a miniature fiber coupled microscope with active axial-scanning. Sci Rep. 2018;8(1):8108. Epub 2018/05/29. doi: 10.1038/s41598-018-26326-3. PubMed PMID: 29802371; PMCID: PMC5970169.
5. Ryan DP, Gould EA, Seedorf GJ, Masihzadeh O, Abman SH, Vijayaraghavan S, Macklin WB, Restrepo D, Shepherd DP. Automatic and adaptive heterogeneous refractive index compensation for light-sheet microscopy. Nat Commun. 2017;8(1):612. doi: 10.1038/s41467-017-00514-7. PubMed PMID: 28931809.
2. Guthman EM, Ma M, Chu P, Baca SM, Restrepo D, Huntsman MM. Cell type specific control of basolateral amygdala plasticity via feedforward inhibition. bioRxiv. 2018. doi: https://doi.org/10.1101/348524.
3. Gould EA, Busquet N, Shepherd D, Dietz RM, Herson PS, Simoes de Souza FM, Li A, George NM, Restrepo D, Macklin WB. Mild myelin disruption elicits early alteration in behavior and proliferation in the subventricular zone. Elife. 2018;7. Epub 2018/02/14. doi: 10.7554/eLife.34783. PubMed PMID: 29436368; PMCID: PMC5828668.
4. Ozbay BN, Futia GL, Ma M, Bright VM, Gopinath JT, Hughes EG, Restrepo D, Gibson EA. Three dimensional two-photon brain imaging in freely moving mice using a miniature fiber coupled microscope with active axial-scanning. Sci Rep. 2018;8(1):8108. Epub 2018/05/29. doi: 10.1038/s41598-018-26326-3. PubMed PMID: 29802371; PMCID: PMC5970169.
5. Ryan DP, Gould EA, Seedorf GJ, Masihzadeh O, Abman SH, Vijayaraghavan S, Macklin WB, Restrepo D, Shepherd DP. Automatic and adaptive heterogeneous refractive index compensation for light-sheet microscopy. Nat Commun. 2017;8(1):612. doi: 10.1038/s41467-017-00514-7. PubMed PMID: 28931809.
Proudly powered by Weebly